The biological phenomena are governed by complex network systems including many species of molecules, cells or organs. For the aim of understanding the functions of complex systems, we adopt mathematical and computational methods.
By theoretical approaches we decipher huge amounts of experimental information, and give integrative understanding for the biological systems.
Our final goal is to open a new bioscience which will progress by the repeats of the theoretical predictions and the experimental verifications.
RESEARCH
-
Complete control of a gene regulatory network of ascidian embryo by a few factors identified by a mathematical theory.
VIEW MORE -
Determining responses of chemical reaction systems from structure of networks
VIEW MORE -
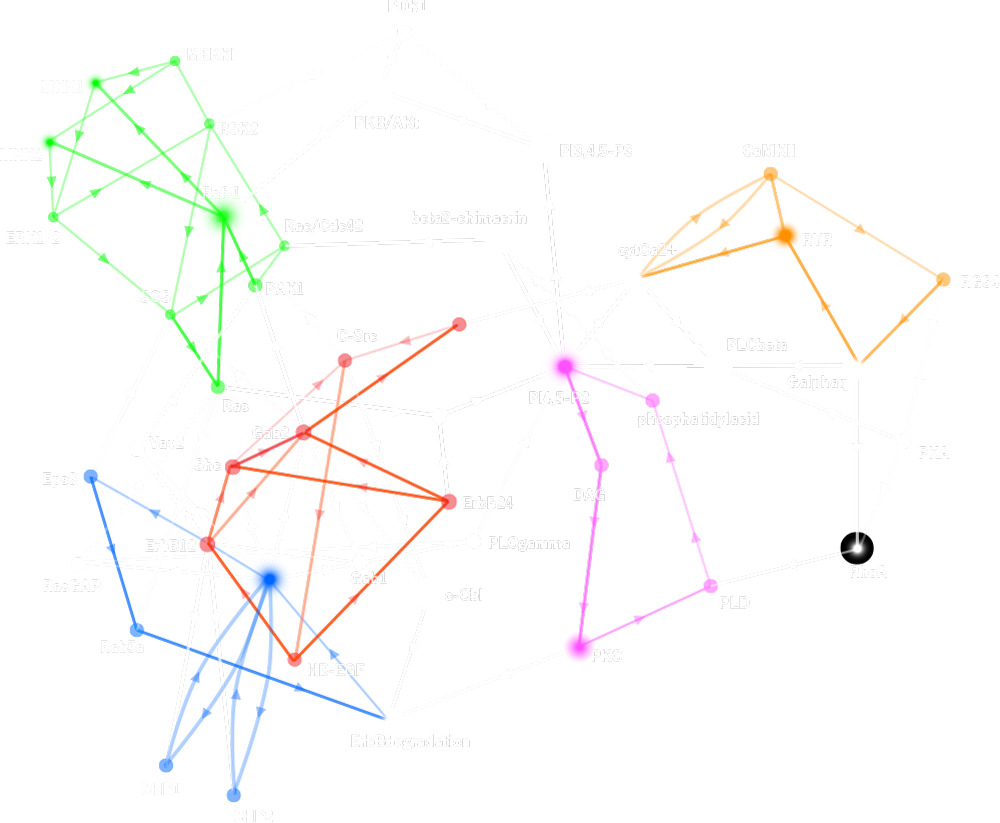
Structure and dynamics of regulatory networks
VIEW MORE -

Physics of organelle morphogenesis
VIEW MORE
NEWS
- publication
Ooka, H., Wintzer, M.E., Komatsu, H., Suda, T., Adachi, K., Li, A., Kong, S., Hashizume, D., Mochizuki, A. and Nakamura, R. (2024) Microkinetic model to rationalize the lifetime of electrocatalysis: trade-off between activity and stability. Journal of Physical Chemistry Letters 15(40), 10079–10085.
- publication
Okino, R., Mukai, K., Oguri, S., Masuda, M., Watanabe, S., Yoneyama, Y., Nagaosa, S., Miyamoto, T., Mochizuki, A., Takahashi, S.-I. and Hakuno, F. (2024) IGF-I concentration determines cell fate by converting signaling dynamics as a bifurcation parameter in L6 myoblasts. Scientific Reports 14, 20699.
- publication
Hosoda, S., Iwata, H., Miura, T., Tanabe, M., Okada, T., Mochizuki, A. and Sato, M. (2024) BayesianSSA: a Bayesian statistical model based on structural sensitivity analysis for predicting responses to enzyme perturbations in metabolic networks. BMC Bioinformatics 25, 297.
- publication
Yu, Q., Ascensao, J.A., Okada, T., COVID-19 Genomics UK (COG-UK) Consortium, Boyd, O., Volz, E. and Hallatschek, O. (2024) Lineage frequency time series reveal elevated levels of genetic drift in SARS-CoV-2 transmission in England. Plos Pathogens, 20(4), p.e1012090.
- publication
Honjo, M., Suzuki, K., Katai, J., Tashiro, Y., Aoyagi, T., Hori, T., Okada, T., Saito, Y. and Futamata, H. (2024) Stable States of a Microbial Community Are Formed by Dynamic Metabolic Networks with Members Functioning to Achieve Both Robustness and Plasticity. Microbes and environments, 39(1), p.ME23091.
- publication
Ooka, H., Wintzer, M.E., Komatsu, H., Suda, T., Adachi, K., Li, A., Kong, S., Hashizume, D., Mochizuki, A. and Nakamura, R. (2024) Microkinetic model to rationalize the lifetime of electrocatalysis: trade-off between activity and stability. Journal of Physical Chemistry Letters 15(40), 10079–10085.
- publication
Okino, R., Mukai, K., Oguri, S., Masuda, M., Watanabe, S., Yoneyama, Y., Nagaosa, S., Miyamoto, T., Mochizuki, A., Takahashi, S.-I. and Hakuno, F. (2024) IGF-I concentration determines cell fate by converting signaling dynamics as a bifurcation parameter in L6 myoblasts. Scientific Reports 14, 20699.
- publication
Hosoda, S., Iwata, H., Miura, T., Tanabe, M., Okada, T., Mochizuki, A. and Sato, M. (2024) BayesianSSA: a Bayesian statistical model based on structural sensitivity analysis for predicting responses to enzyme perturbations in metabolic networks. BMC Bioinformatics 25, 297.
- publication
Yu, Q., Ascensao, J.A., Okada, T., COVID-19 Genomics UK (COG-UK) Consortium, Boyd, O., Volz, E. and Hallatschek, O. (2024) Lineage frequency time series reveal elevated levels of genetic drift in SARS-CoV-2 transmission in England. Plos Pathogens, 20(4), p.e1012090.
- publication
Honjo, M., Suzuki, K., Katai, J., Tashiro, Y., Aoyagi, T., Hori, T., Okada, T., Saito, Y. and Futamata, H. (2024) Stable States of a Microbial Community Are Formed by Dynamic Metabolic Networks with Members Functioning to Achieve Both Robustness and Plasticity. Microbes and environments, 39(1), p.ME23091.